Codelet Extractor and REplayer
Introducing CERE
What is CERE?
Codelet Extractor and REplayer (CERE) is an open source framework for code isolation developped jointly at UVSQ Li-PaRAD lab and ECR lab. CERE finds and extracts the hotspots of an application as isolated fragments of code, called codelets. Codelets can be modified, compiled, run, and measured independently from the original application. Code isolation reduces benchmarking cost and allows piecewise optimization of an application. Unlike previous approaches, CERE isolates codes at the compiler Intermediate Representation level. Therefore CERE is language agnostic and supports many input languages such as C, C++, Fortran, and D. CERE automatically detects codelets invocation that have the same performance behavior. Then, it selects a reduced set of representative codelets and invocations, much faster to replay, which still captures accurately the original application. In addition, CERE supports recompiling and retargeting the extracted codelets. Therefore, CERE can be used for cross-architecture performance prediction or piecewise code optimization. On the SPEC 2006 FP benchmarks, CERE codelets cover 90.9% and accurately replay 66.3% of the execution time. We use CERE codelets in a realistic study to evaluate three different architectures on the NAS benchmarks. CERE accurately estimates each architecture performance and is 7.3 times to 46.6 times cheaper than running the full benchmark. Within the Montblanc European Research Project CERE is being ported to ARM Aarch64 architectures.
Publications
CERE: LLVM based codelet extractor and REplayer for piecewise benchmarking and optimization. Pablo de Oliveira Castro, Chadi Akel, Eric Petit, Mihail Popov, and Jalby William. ACM Transactions on Architecture and Code Optimization, 2015. [ bib | DOI ]
PCERE: Fine-grained Parallel Benchmark Decomposition for Scalability Prediction. Mihail Popov, Chadi Akel, Florent Conti, William Jalby, and Pablo de Oliveira Castro. In Proceedings of the 29th IEEE International Parallel and Distributed Processing Symposium IPDPS 2015. IEEE, 2015. [ bib ]
Piecewise Holistic Autotuning of Compiler and Runtime Parameters. Mihail Popov, Chadi Akel, William Jalby, and Pablo de Oliveira Castro. In Euro-Par 2016 Parallel Processing - 22nd International Conference, Lecture Notes in Computer Science. Springer, 2016. [ bib ]
Automatic Decomposition of Parallel Programs for Optimization and Performance Prediction. Mihail Popov, PhD thesis, defended in 2016.
Piecewise Holistic Autotuning of Parallel Programs with CERE. Mihail Popov, Chadi Akel, Yohan Chatelain, William Jalby, and Pablo de Oliveira Castro. Concurrency and Computation: Practice and Experience, 2017. [ bib ]
Efficient Thread/Page/Parallelism Autotuning for NUMA Systems. Mihail Popov, Alexandra Jimborean, and David Black-Schaffer. International Conference on Supercomputing (ICS '19). ACM, 2019. [ bib ]
How does it works?
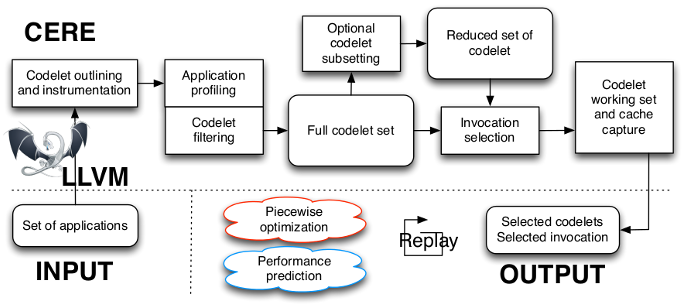
CERE takes as an input the source files of an application or a benchmark suite. All the languages supported by the LLVM front-ends (C, C++, Fortran, D, etc.) are accepted. The application loops are outlined and instrumented with profiling probes to identify the application hotspots. The hot loops are kept and form the full codelet set. CERE can prune this full codelet set and only keep a reduced set of representative codelets. Then, a clustering algorithm analyzes the invocation performance trace of each codelet to find a representative subset of invocations. The memory and cache state of each representative invocation is then captured and dumped to disk. The output of this process is a set of representative codelets and invocations, which can be redistributed, recompiled and replayed on different systems and architectures. The codelet set can be used as a proxy for original application in optimization or benchmarking studies.
Get in touch
For questions or discussions please use the cere-dev@googlegroups.com mailing list.
Getting CERE
Download CERE
CERE is distributed under the GNU Lesser General Public License. Sources and latest releases are available at https://github.com/benchmark-subsetting/cere
Install CERE
Please follow the instructions here.
Replay Accuracy Reports
NAS 3.0 SER
| Benchmark | Description |
|---|---|
| BT | Block Tri-diagonal solver |
| CG | Conjugate Gradient |
| EP | Embarrassingly Parallel |
| FT | Discrete 3D fast Fourier Transformation |
| IS | Integer Sort |
| LU | Lower-Upper Gauss-Seidel solver |
| MG | Multi-Grid on a sequence of meshes |
| SP | Scalar Penta-diagonal solver |
SPEC CPU 2006
| Benchmark | Description |
|---|---|
| 410-bwaves | Fluid Dynamics |
| 416-gamess | Quantum Chemistry |
| 433-milc | Quantum Chromodynamics |
| 434-zeusmp | CFD |
| 435-gromacs | Biochemistry: Molecular Dynamics |
| 436-cactusADM.html | General Relativity |
| 437-leslie3d | Fluid Dynamics |
| 444-namd | Biology: Molecular Dynamics |
| 447-dealII | Finite Element Analysis |
| 450-soplex | Linear Programming, Optimization |
| 453-povray | Image Ray-tracing |
| 454-calculix | Structural Mechanics |
| 459-GemsFDTD | Computational Electromagnetics |
| 465-tonto | Quantum Chemistry |
| 470-lbm | Fluid Dynamics |
| 481-wrf | Weather Prediction |
| 482-sphinx3 | Speech recognition |
| 999-specrand | Random Number Generation |
| Benchmark | Description |
|---|---|
| lulesh2.0 Intel Haswell | Hydrodynamics |
| lulesh2.0 Aarch64 | Hydrodynamics |